publications
2026
-
 Convex estimation of Gaussian graphical regression models with covariatesRuobin Liu and Guo Yusubmitted, 2026
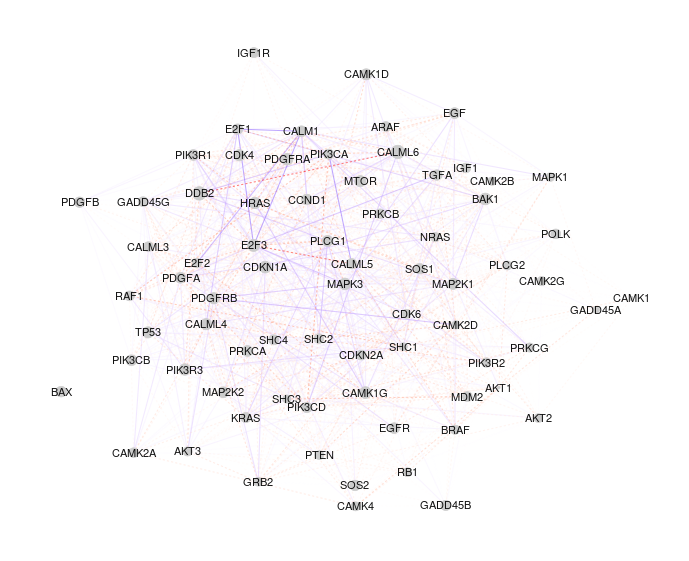
Convex estimation of Gaussian graphical regression models with covariatesRuobin Liu and Guo Yusubmitted, 2026Gaussian graphical models (GGMs) are widely used to recover the conditional independence structure among random variables. Recent work has sought to incorporate auxiliary covariates to improve estimation, particularly in applications such as co-expression quantitative trait locus (eQTL) studies, where both gene expression levels and their conditional dependence structure may be influenced by genetic variants. Existing approaches to covariate-adjusted GGMs either restrict covariate effects to the mean structure or lead to nonconvex formulations when jointly estimating the mean and precision matrix. In this paper, we propose a convex framework that simultaneously estimates the covariate-adjusted mean and precision matrix via a natural parametrization of the multivariate Gaussian likelihood. The resulting formulation enables joint convex optimization and yields improved theoretical guarantees under high-dimensional scaling, where the sparsity and dimension of covariates grow with the sample size. We support our theoretical findings with numerical simulations and demonstrate the practical utility of the proposed method through a reanalysis of an eQTL study of glioblastoma multiforme (GBM), an aggressive form of brain cancer.
2025
-
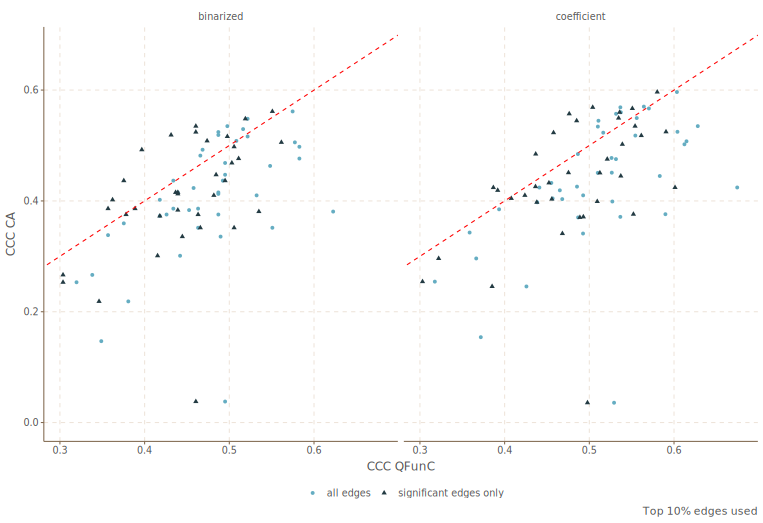 A Mixed Model Approach for Estimating Regional Functional Connectivity from Voxel-level BOLD SignalsRuobin Liu, Chao Zhang, Chau Tran, Sophie Achard, Wendy Meiring, and Alexander Petersensubmitted, 2025
A Mixed Model Approach for Estimating Regional Functional Connectivity from Voxel-level BOLD SignalsRuobin Liu, Chao Zhang, Chau Tran, Sophie Achard, Wendy Meiring, and Alexander Petersensubmitted, 2025Resting-state brain functional connectivity quantifies the synchrony between activity patterns of different brain regions. In functional magnetic resonance imaging (fMRI), each region comprises a set of spatially contiguous voxels at which blood-oxygen-level-dependent signals are acquired. The ubiquitous Correlation of Averages (CA) estimator, and other similar metrics, are computed from spatially aggregated signals within each region, and remain the quantifications of inter-regional connectivity most used by neuroscientists despite their bias that stems from intra-regional correlation and measurement error. We leverage the framework of linear mixed-effects models to isolate different sources of variability in the voxel-level signals, including both inter-regional and intra-regional correlation and measurement error. A novel computational pipeline, focused on subject-level inter-regional correlation parameters of interest, is developed to address the challenges of applying maximum (or restricted maximum) likelihood estimation to such structured, high-dimensional spatiotemporal data. Simulation results demonstrate the reliability of correlation estimates and their large sample standard error approximations, and their superiority relative to CA. The proposed method is applied to two public fMRI data sets. First, we analyze scans of a dead rat to assess false positive performance when connectivity is absent. Second, individual human brain networks are constructed for subjects from a Human Connectome Project test-retest database. Concordance between inter-regional correlation estimates for test-retest scans of the same subject are shown to be higher for the proposed method relative to CA.
-
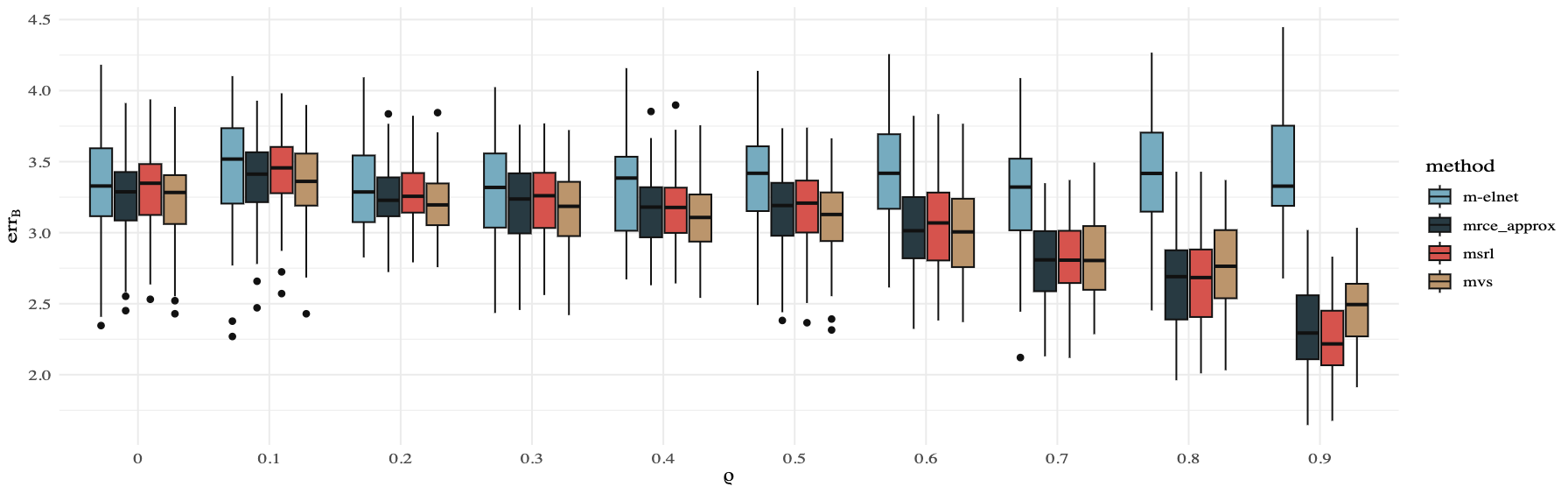 Estimation of the Error Structure in Multivariate Response Linear Regression ModelsRuobin Liu and Guo YuWIREs Computational Statistics, 2025e70021 EOCS-730.R1
Estimation of the Error Structure in Multivariate Response Linear Regression ModelsRuobin Liu and Guo YuWIREs Computational Statistics, 2025e70021 EOCS-730.R1Multivariate response linear regression model considers how a set of covariates affects multiple responses. In contrast to separately running univariate regression for each response, multivariate response regression can better estimate the coefficient matrix by exploiting shared information among the responses. A key to effectively borrowing strength across different responses is to estimate and utilize the structure among random errors in responses. Moreover, certain applications seek to understand the interrelationships of the responses, which are directly encoded in the error structure. These observations call for the development of methods for estimating the error structure in multivariate response linear regression models. This article aims at providing a review of recent progress in this challenging and important problem. We also provide simulation studies demonstrating the empirical performance of recently developed methods for error structure estimation, as well as the benefit by exploiting these estimates for regression coefficients estimation.
2024
-
 Deep residual networks for crystallography trained on synthetic dataDerek Mendez, James M. Holton, Artem Y. Lyubimov, Sabine Hollatz, Irimpan I. Mathews, Aleksander Cichosz, Vardan Martirosyan, Teo Zeng, Ryan Stofer, Ruobin Liu, Jinhu Song, Scott McPhillips, Mike Soltis, and Aina E. CohenActa Crystallographica Section D, Jan 2024
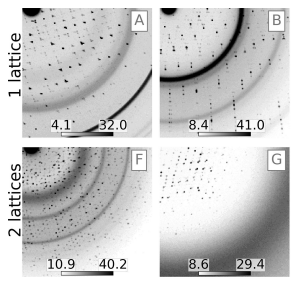
Deep residual networks for crystallography trained on synthetic dataDerek Mendez, James M. Holton, Artem Y. Lyubimov, Sabine Hollatz, Irimpan I. Mathews, Aleksander Cichosz, Vardan Martirosyan, Teo Zeng, Ryan Stofer, Ruobin Liu, Jinhu Song, Scott McPhillips, Mike Soltis, and Aina E. CohenActa Crystallographica Section D, Jan 2024The use of artificial intelligence to process diffraction images is challenged by the need to assemble large and precisely designed training data sets. To address this, a codebase called \it Resonet was developed for synthesizing diffraction data and training residual neural networks on these data. Here, two per-pattern capabilities of \it Resonet are demonstrated: (i) interpretation of crystal resolution and (ii) identification of overlapping lattices. \it Resonet was tested across a compilation of diffraction images from synchrotron experiments and X-ray free-electron laser experiments. Crucially, these models readily execute on graphics processing units and can thus significantly outperform conventional algorithms. While \it Resonet is currently utilized to provide real-time feedback for macromolecular crystallography users at the Stanford Synchrotron Radiation Lightsource, its simple Python-based interface makes it easy to embed in other processing frameworks. This work highlights the utility of physics-based simulation for training deep neural networks and lays the groundwork for the development of additional models to enhance diffraction collection and analysis.